 Anaheim Convention Center |
 EB2010 EXPERIMENTAL BIOLOGY MEETINGS Anaheim, CA APRIL 24-28, 2010  |
 Anaheim Convention Center |
 EB2010 EXPERIMENTAL BIOLOGY MEETINGS Anaheim, CA APRIL 24-28, 2010  |
The University of Delaware group included four faculty and 13 undergraduates.
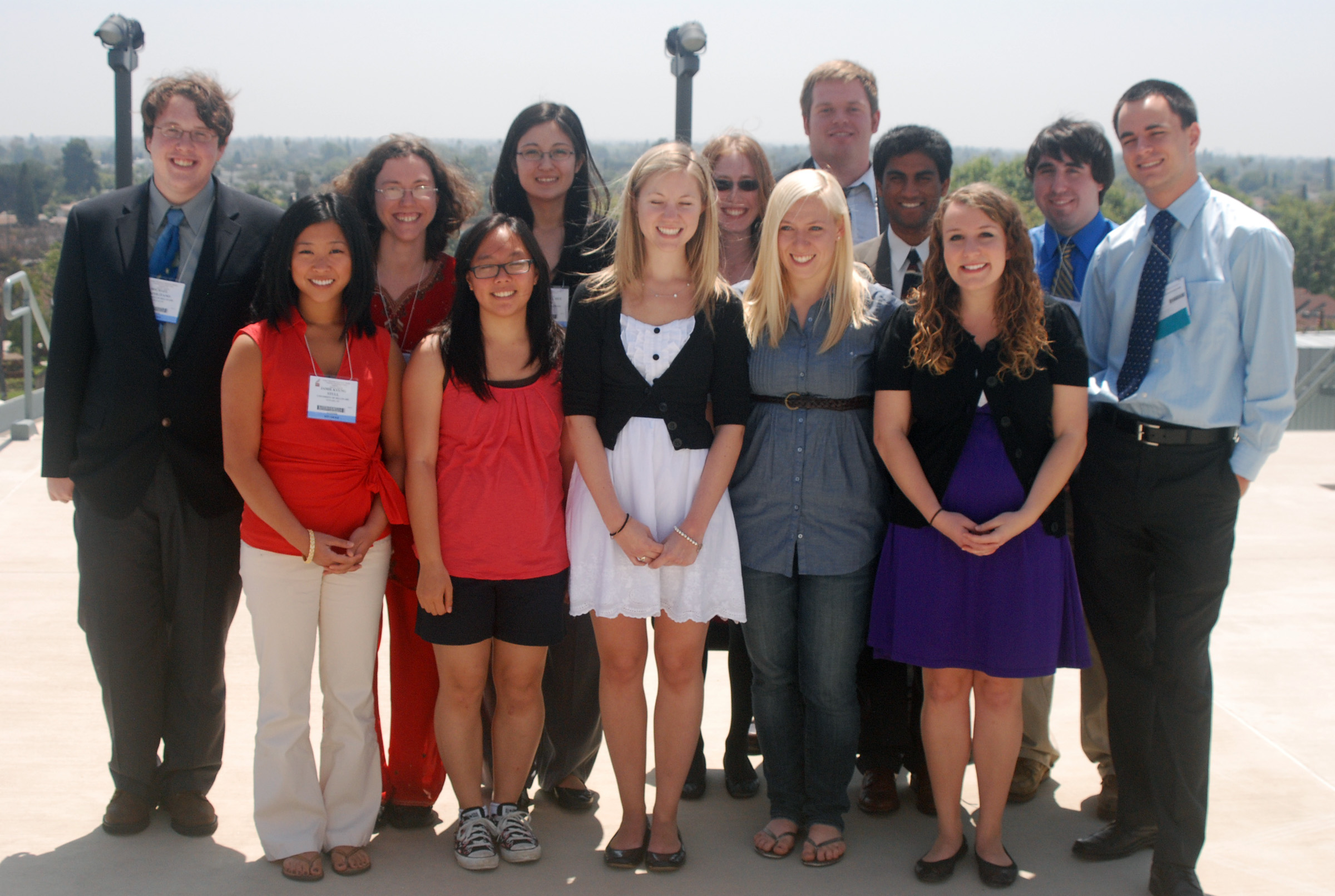 From left to right: Michael Napolitano, Jamie Stull, Amy Styer, Jean Huynh, Rebecca Brown, Megan Kissig, Laura Sloofman, Katharine Shelly, Tyler Larsen, Tejal Naik, Rachel Randell, Robert Sheehan, and Aleksey Dvorzhinskiy. |
||||
| Prof.
Hal White, Chem & Biochem Prof. Dave Usher, Biol. Sci. Prof. Seung Hong, Biol Sci Prof. Gary Laverty, Biol Sci |
Rebecca Brown Aleksey Dvorzhinskiy Jean Huynh Megan Kissig |
Tyler
Larsen Tejal Naik Michael Napolitano Wachen Peters |
Rachel Randell Robert Sheehan Katharine Shelly |
Laura
Sloofman Jamie Stull Amy Styer |
|
|
Rebecca
Sul Hee Brown1, Dong-Hoon Jeong2,3,
Blake C Meyers2,3, Pamela J Green2,3
One
way plants respond to environmental stress is by modifying their gene
expression through the use of microRNAs (miRNAs). miRNAs are
noncoding small RNAs that regulate
gene expression at the posttranscriptional level by base pairing with
complementary messenger RNA (mRNA) molecules, causing either mRNA
cleavage or
translational inhibition. To elucidate
the roles of miRNAs in environmental stress responses, wild type plants
and
miRNA enriched mutant plants were subjected to various environmental
stresses. Small RNA libraries from
stress-treated seedlings and flowers were constructed and sequenced
using deep
sequencing technologies. Computational
analysis revealed differential expression of miRNAs between
stress-treated
plants and control plants with some miRNAs upregulated and some
downreguled by
environmental stress. Additional
analysis revealed potentially novel miRNAs in Arabidopsis as
well.
These results suggest that miRNAs play
important roles in stress responses, and that miRNA identification in
Arabidopsis has not been saturated.
Future work will involve validating highly stress-regulated miRNAs and
prospective new miRNAs, and examining their potential target genes for
regulation. Stress-regulated miRNAs will
be incorporated into multi-network models and their biological roles
will be
hypothesized and tested through functional studies. R.S.H.B. was
supported by the Howard Hughes
Medical Institute, and NSF grant MCB#0548569 to P.J.G. provided
research
support. Abstract Number: 4114 |
 Aleksey Dvorzhinskiy Recipient of an ASBMB Undergraduate Travel Award.
|
Abstract number: 2838 |
|
Mammalian sperm are dependent on JAM-A for normal motility: Jean
Huynh, Rolands Aravindan, and Patricia
A. Martin-DeLeon Defective
sperm
motility (asthenozoospermia) is a primary cause of male infertility.
Junctional
Adhesion Molecule-A (JAM-A) has been recently shown to be essential for
mouse sperm
motility. However, to date it has not been studied in human sperm. The
objective of this study is to determine if JAM-A is present in human
sperm and to
characterize its expression. In silico
analysis reveals 83% similarity in the amino acid sequence of JAM-A in
mice and
humans, indicating that the proteins are conserved. This suggests
that JAM-A may also play a role in
human sperm motility. Western analysis of human sperm proteins revealed
the
presence of JAM-A with a MW of ~32kDa, identical to that of the mouse.
Immunocytochemistry
was used to localize the expression of JAM-A in human sperm. Flow
cytometry
confirmed the Western blot data and revealed the presence of the
protein on the
sperm membrane in one of two individuals attending an infertility
clinic. Once
JAM-A has been fully characterized in human sperm, the next step will
be to determine
its interacting protein partners by attempting to co-immunoprecipitate
it with candidate
proteins. The localization and characterization of JAM-A in human sperm
is
expected to increase our understanding of genetic factors leading to
human male
infertility and subfertility, laying the groundwork for diagnosis of a
subset
of couples with fertility issues. Funded by NIH-COBRE grant
#5P20RR015588-07. Abstract Number: 6236 |
|
Megan Kissig
|
Megan E. Kissig and Ulhas P. Naik
Department of Biological Sciences Non-alcoholic
fatty liver disease (NAFLD) is characterized by an abnormal amount of
fat accumulation
in the liver, specifically more than 5% fat by weight. Little is known
about
how the fat accumulates in the liver, but it has been found that
intestinal permeability
due to leaky tight junctions may be a contributing factor. Our lab
studies junctional
adhesion molecule-A (JAM-A), a protein located at the tight junctions
of epithelial
and endothelial cells. Through its ability to homodimerize at the
apical part
of the lateral membrane, JAM-A helps regulate the permeability and
stability of
the junction. This study’s aim was to find what effect JAM-A has on the
development of NAFLD. To analyze this relationship, groups of Jam-A (+/+) and Jam-A (-/-) mice were
put on either a high-fat or low-fat diet for
20 weeks. During this time the mice were weighed every two weeks and
blood
samples were taken every four weeks. At the end of the 20 weeks, the
mice were
sacrificed and the livers and fat pads were removed. We found that the
variation
in weight between the Jam-A (-/-)
groups was greater than that between the Jam-A
(+/+) groups. Also, the average area of the adipocytes in the high-fat Jam-A (-/-) group was greater than the
average area in the high-fat Jam-A
(+/+) group. Since the liver sends fat to tissues in the form of
cholesterol,
we compared the levels in the plasma. The high-fat Jam-A (-/-)
mice had significantly higher LDL-cholesterol and total
cholesterol levels in the plasma than the other groups as well. After
examining
the stained sections of the livers, it
was found that the livers of the high-fat Jam-A (-/-)
mice showed more fat droplet accumulation than the
high-fat Jam-A (+/+) mice. This suggests that
ultimately
the absence of JAM-A from the tight junctions promotes the development
of
NAFLD. This project was funded by the Arnold and Mabel Beckman
Foundation.
|
 Tyler Larsen |
Inherantly
Antibacterial Hydrogels –
Altering ACctivity Via Tryptophan/Arginine Iinteractions Tyler Larsen, Daphne A. Salick, Radhika Nagarkar, Joel P. Schneider Department of Chemistry and Biochemistry, University of Delaware, Newark, DE 19716 Hydrogels are heavily
hydrated materials that show considerable promise
as artificial extracellular matrices for use in tissue regenerative
therapies. The development of antibacterial hydrogels has been of great
interest to the hydrogel research community as a means to combat the
threat of infection during material implantation. We have
developed MAX1, a self-assembling b-hairpin peptide hydrogel whose
surface exhibits inherent antibacterial activity against several
pathogens prevalent in hospital settings. Under physiological
conditions, MAX1 self-assembles into a crosslinked, mechanically rigid
hydrogel, but the process is too slow for use as an injectable
implant. This study aims to design a hydrogel with rapid
self-assembly and high rigidity under physiological conditions while
maintaining potent inherent antibacterial activity through
incorporation of cation-p interactions, which are common in
antibacterial peptides found in nature. A new b-hairpin peptide
(RWMAX1) was designed, incorporating a cross-strand R/W pair into the
MAX1 sequence. The folding and self-assembly properties were
assessed using circular dichroism and rheology and the antibacterial
activity was investigated against gram positive S. aureus and gram
negative E. coli.
RWMAX1 gels were found to possess both
favorable physical and antimicrobial properties. Supported
by an Undergraduate Beckman
Award.
Abstract Number
2386 |
 Tejal Naik ASBMB 2010 Thematic Best Poster selected by the theme organizers for its outstanding research in Chemical Biology and Drug Discovery. |
Development of a Peptide Nucleic Acid Based siRNA Delivery System Tejal U. Naik and Millicent O. Sullivan Small interfering RNA (siRNA) plays a major role in gene silencing. The ability to harness siRNA for therapeutic benefit can have a widespread impact on a variety of diseases. However, the efficacy of such treatments is limited due to the many barriers associated with siRNA delivery. In this work, we lay the groundwork for the development of a peptide nucleic acid (PNA)-based surface-mediated siRNA delivery system. PNAs are nucleic acid analogs that hybridize with complementary DNA or RNA sequences, enabling the direct attachment of various macromolecules such as peptides. Conjugation of targeting and protective moieties can potentially enhance the delivery of siRNA. We first established a cell transfection model utilizing stably transfected B16FO mouse melanoma cells producing GFP. Anti-GFP siRNA was designed and its efficacy was evaluated via flow cytometry and fluorescence microscopy. Optimization of cell seeding density, siRNA concentration, use of antibiotics, and time of transfections was accomplished. Our second task was to prepare molecular conjugates for PNA-based siRNA modifications. PNA-peptide conjugates were assembled and purified through Reverse Phase HPLC. Future development of this delivery system will include linkage of anti-GFP siRNA to surfaces via the PNA-peptide tethers, and exploration of the delivery system in B16FO cells.Abstract Number: 6946 |
 Michael Napolitano |
of Vibrio Pathogenicity
Island 2
Michael
G. Napolitano, Moreno, S. A., Duncan, M.,
and Boyd,
E. F. Department of Biological Sciences Abstract Number: 1976
|
 Wachen Peters |
P2 Receptors and ATP Enhance the Migration of Prostate Cancer Cells
Wachen
Peters, Christine Maguire2, Randall
L. Duncan2, and Robert A.
Sikes2Lincoln
University1
and University of Delaware2 Purinergic
signaling stimulates many biological processes. Two classes of
purinergic
receptors, GPCR (P2Y) and gated channels (P2X), have been identified
that bind
ATP as a ligand, which can promote cell migration, a critical component
of metastasis.
ATP also has been shown to have an anticancer effect in vivo. The
purpose of
this study was to ascertain the effect of ATP on the migration of an
isogenic
progression series of prostate cancer (PCa) cell lines on three
different
extracellular matrices. We hypothesize that ATP treatment activates
purinergic
receptors that increases cell migration on all three extracellular
matrices
(ECM).
Funded
by: Department
of Defense HBCU/MI Undergraduate Research Training Grant, PC080950; NIH
INBRE
P20RR016472, NIH COBRE P20
RR015588 Abstract Number: 1053 |
|
|
Heparan
and chondroitin sulfate modifications in signaling pathways
regulating articular cartilage homeostasis
Rachel Leigh Randell, Richard Wittmeyer, and Erica M. Selva Degradation of articular cartilage leads to osteoarthritis (OA) in humans. Articular cartilage homeostasis depends on signaling events modulated by extracellular matrix glycoproteins. We tested the hypothesis that disrupting heparan sulfate (HS) and chondroitin sulfate (CS) proteoglycans would alter signaling pathways relevant to cartilage homeostasis. Mosaic mutant and wild-type Drosophila wing discs were used to visualize the effects of mutations in two genes: O-xylosyltransferase (oxt), required for linking HS and CS to core proteins, and sulfateless (sfl), essential for decorating HS with negatively charged sulfate groups. We performed immunostaining of signaling ligands and pathway activation targets, including: Decapentaplegic, the homolog of human Bone Morphogenic Protein involved in anabolic cartilage processes; Wingless, the homolog of human Wnts implicated in cartilage repair; and Hedgehog and its receptor Patched. We demonstrate the importance of HS and CS for proper activation and movement of signaling molecules. Surprisingly, sfl appears to be more disruptive than oxt on signaling. Ongoing experiments examine the role of the core proteoglycan Dally-like and look for signals that might upregulate cartilage deposition. These results will help to identify targets for the development of novel OA therapeutics. Supported in part by an HHMI undergraduate research award. Abstract Number: 6312 |
|
The
Effects of Histone Modifications on Lens Cell Denucleation Robert
Patrick Sheehan and Melinda K.
Duncan
Department of Biological Sciences, University of Delaware, Newark, DE, 19716 The
lens of the eye contains
both epithelial and fiber cells, with fibers cells deriving from the
equatorial
epithelium. When these cells differentiate, the nucleus and cellular
organelles
are broken down to facilitate transparency. However, small fragments of
DNA can
remain in fully mature lens fiber cells although its structure is
unknown. We found
that histone H3, trimethylated on Lysine 9, which is associated with
compact
heterochromatin, co-localizes with these DNA fragments.
In contrast, the DNA remnants are not
associated with either histone H3 Lysine 9 acetylation or histone H4
lysine 8
acetylation which is predominant in the more accessible euchromatin.
These data
suggest that the DNA fragments persisting in mature lens fiber cells
are
entirely composed of heterochromatin, perhaps because their highly
compacted
state prevented access of the nucleases responsible for DNA degradation
during
lens fiber cell differentiation. Supported
by Howard Hughes Medical Institute and the National Eye Institute. Abstract
Number: 6387 |
|
Localization of human
sperm PMCA4b which is required for normal motility in mouse sperm
Katharine E Shelly, Rolands Aravindan, and Patricia A Martin-DeLeon Department of Biological Sciences Progressive and hyperactivated motility in
sperm are both dependent on Ca2+ and ATP to effect
fertilization, with
hyperactivated motility utilizing more ATP than progressive motility. The ATP-driven enzyme Plasma Membrane
Calcium/calmodulin-dependent ATPase isoform 4b
(PMCA4b) is
the principal calcium efflux pump in mouse sperm, and its absence
results in
the loss of hyperactivated motility and fertility.
Thus, PMCA4b plays an important role in
murine sperm function. Protein BLAST
analysis illustrates 84% identity and 91% similarity between the mouse
and
human PMCA4b sequences. Analysis of the
functional domains suggests conservation of PMCA4b’s activity in human
sperm. The objective of this
investigation is to characterize the presence of PMCA4b in human sperm
by
Western blotting, as well as to determine its subcellular localization
by
immunocytochemistry (ICC). Flow
cytometric analysis serves to confirm the presence of PMCA4b on the
membrane.
ICC studies should elucidate whether or not localization of human
PMCA4b is present
on the principal piece of the flagellum, which would coincide with
findings in
the mouse and suggest its involvement in human sperm motility. This
work was
supported by the Charles Peter White Fellowship and the NIH-COBRE grant
#5P20RR015588-07. Abstract Number: 6308 |
 Laura Sloofman Recipient of an Honorable Mention Award in the ASBMB Undergraduate Poster Competition. |
Mathematical
modeling of elasticity changes in the
cortical bone Mathematical
modeling, or using equations to describe a system, is a powerful
technique in
transgenic research. Our laboratory created the first viable mutant
mouse model
lacking an individual ribosomal protein (HIP/RPL29). A
short stature phenotype associated with increased bone fragility is
observed in
HIP/RPL29 null mice. We recently noted that a decrease in collagen
cross-linking during the growth of HIP/RPL29 null bone precedes an
overall
enhancement in the mineral-to-matrix ratio in adult bone. We
hypothesize
that we can build on existing mathematical models to describe the
changes in
bone structure and composition resulting from HIP/RPL29 deficiency.
Hierarchical multi-scale models of wild type and HIP/RPL29 null
cortical bone
will be used to simulate adult control and mutant bone. We aim to
relate the
changes in null bone samples to elasticity measurements collected via
three-point bending tests. Since the equations describing mutant and
control
bone are interrelated, we can extrapolate relationships among the
mentioned
properties. We will also use this model to predict the elasticity in
control
and mutant bones at three months of age. All calculations will be
compared with
accepted experimental values. This model will be altered as the
three-month
mutant bone phenotype is refined. Funding was provided by the Howard
Hughes
Medical Institute to LGS and NIH P20-RR016458 to CBKS. Abstract
Number 5191 |
|
Lenses from
Connexin50 Mutant Mice Exhibit
the Unfolded Protein Response Jaime K. Stull, Zeynep Firtina,
and Melinda K. Duncan Cataract is the leading cause of blindness worldwide. Although the majority of cataracts are age-related, they may also be caused by heritable mutations. In humans as well as mice, congenital cataract occurs in Connexin50 (Cx50) mutants although the cataract phenotypes are more severe than that observed in Cx50 null mice. We hypothesized that when cells attempt to make Cx50 from a mutated gene, the resulting protein does not fold properly, inducing a cell-stress mechanism called the Unfolded Protein Response, UPR. We investigated this in mice harboring two different Cx50 mutations, Cx50S50P/S50P and Cx50G22R/G22R, that both exhibit similar lens phenotypes including microphthalmia, unorganized fiber cell structure, and cataract. We found a noticeable upregulation of the expression of the molecular folding chaperone BiP in mutant lenses after E15.5 in addition to an upregulation of XBP-1 gene expression and unconventional splicing indicative of the activation of the UPR sensor IRE1. These data further support our prior work suggesting that UPR can contribute to lens phenotypes due to mutations in genes whose products must transit the secretory pathway. Supported in part by an HHMI undergraduate research award. |
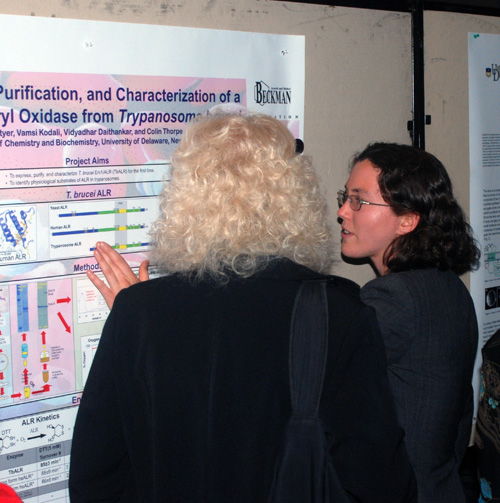 Amy Styer Recipient of an Honorable Mention Award in the ASBMB Undergraduate Poster Competition. Recipient of an ASBMB Undergraduate Travel Award.
|
Amy Styer, Vamsi Kodali,
Vidyadhar
Daithankar, and Colin Thorpe
Department of Chemistry and Biochemistry A
sulfhydryl oxidase from Trypanosoma brucei was expressed, purified and
studied enzymologically for the first time.
The protein is homologous to ALR (augmenter of liver regeneration), an
essential flavin-linked enzyme which catalyzes disulfide bond formation
in the
mitochondrial intermembrane space (IMS) of eukaryotes.
Trypanosomes lack a gene for Mia40, a
necessary redox partner with ALR in yeast and mammals. Future
research will determine how
trypanosomes compensate for the lack of Mia40, and if this sulfhydryl
oxidase
has the same biological localization and function as ALR. We have shown
that six
of seven cysteines in the 33 kDa trypanosomal ALR are present as
disulfide
bonds. In oxygen electrode assays, the
enzyme catalyzed disulfide-bond formation in the model substrate
dithiothreitol
(DTT), but not in the monothiols glutathione and cysteine.
Molecular oxygen was a much better terminal
electron acceptor for trypanosome ALR than for human ALR (TbALR
Km = 15±1μM; some 15-fold lower than the human
enzyme). Further studies are aimed at
characterizing the kinetic mechanism of this flavoenzyme by stopped
flow
spectrophotometry. Understanding trypanosome redox biochemistry
is important because Trypanosoma brucei, the causative agent of African
Sleeping sickness, kills nearly fifty thousand people annually and
current
treatments are expensive and incur serious side reactions. This
exploration of trypanosome
mitochondrial IMS disulfide-bond formation may lead to biomedical
advances in
the fight against trypanosome diseases.
Funded by a Beckman Scholars Fellowship to A.S. and NIH GM26643 to C.T. |
  Laura Sloofman
displays her Honorable Mention Award from the ASBMB Undergraduate
Poster Competition in the Systems Biology Category.
|
 Amy Styer displays
her Honorable Mention Award from the ASBMB Undergraduate Poster
Competition in the Protein Structure and Function Category.
|
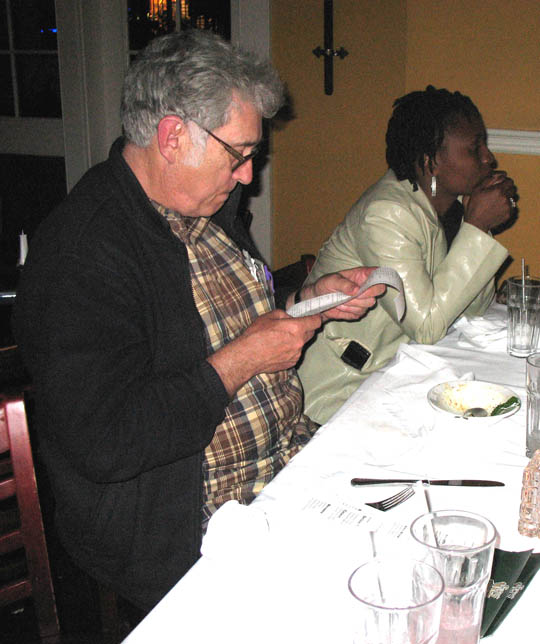 Dr. White wonders how
he will be able to pay for all of the big appetites.
|
 Our Brief excursion into Disneyland. |
 |
 Judge Michael Cox (UD BA Biology '74) at Tejal's poster. |
 Amy, Michael, Rebecca, Seung, and Gary |
 Tejal, Megan, Jean, Kate, and Rachel |
 Tyler, Seung, and Amy |
 Yes, there really are palm trees in Southern California |
Photos by David Usher and Hal White |
 Laura And Megan |
The trip to the Experimental Biology
2010 Meetings
in Anaheim was organized by the University of Delaware HHMI
Undergraduate
Science Education Program with additional support from travel grants
from
the American
Society for Biochemistry and Molecular Biology,
Arnold
and Mabel Beckman
Scholars Program, and the Women
Scholars Program. The HHMI
Undergradaute Science Education Program, the Arnold
and Mabel Beckman
Scholars Program, Charles Peter White
Fund, Undergraduate Research
Program, NIH, NSF, National Eye Institute, and DOD supported
research by the students.